NA-plot manual
This program reads the results of program nab and plots various Bayesian measures of information (marginals, posterior covariance matrices, resolution kernels etc) calculated by nab and written to the file nab.out.
It will read up to two nab.out files and plot the results next to each other. The usual situation would be that one file is the result of running nab on a posterior and the other on a prior.
Type of plot:
The program naplot reads the file `nab.out' produced by the program nab and draws seven types of plot in either X (naplot-x) or Postscript (naplot-p). If Postscript is used then a file called 'naplot.ps' is produced. The major features of each plot are controlled by command line arguments. The minor features are controlled by an input file called `naplot.cmd' (read in from subdirectory `plot'). Below is the syntax for each type of plot and a set of examples that can be run on the demo results files provided in the package.
Plotting 1-D marginals of the posterior probability distribution (PPD)
Plotting 2-D marginals of the posterior probability distribution (PPD)
Plotting the convergence of integrals as a function of sample size
Input files:
nab_post - file name read in from command line (see examples) and contains nab output file for posterior. (must be present)
nab_prior - file name read in from command line (see examples) and contains nab output file for prior. (Optional file)
pal.in - defines colour of pens in RGB (must be present). (See example file for explanation of entries, and examples below.)
model.ref - defines a reference model to be plotted on marginals and convergence plot. (Optional file).
confidence_cont - percentage confidence contours to plot on top of 2-D marginal distributions. (Optional file) (See example file for explanation of entries, and examples below.)
inv_prior_cov.matrix - contains inverse prior covariance matrix. (Optional file). This is used only if no prior input file is specified on the command line. It is usually used when the prior covariance matrix is simple and its inverse is known analytically.
naplot.cmd - contains various plot options to change appearance of plots (optional file). If it is not present then defaults are used. (See example file for explanation of entries.)
finetune - contains increments for moving axis labels and tick marks for a postscript plot (i.e. finetuning). (Optional file). (See example file for explanation of entries.)
Output files:
correlation.matrix - contains values of correlation matrix calculated and plotted by naplot.
resolution.matrix - contains values of resolution matrix calculated and plotted by naplot.
All files, except nab_post and nab_prior , are either read from, or written to, the subdirectory plot
Plotting 1-D marginals of the posterior probability distribution (PPD)
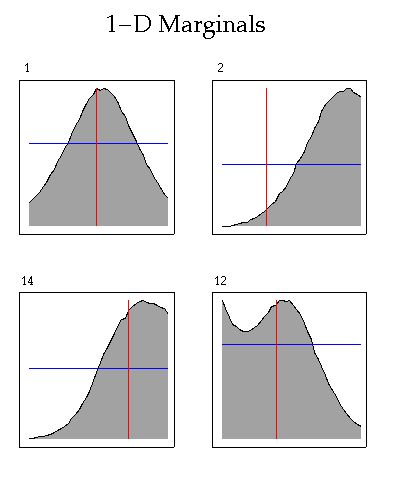
Calling sequence
naplot-x plottype n filename_post (filename_prior) (-r) (-f)
plottype = `1dmarg' to activate.
n = number of plots to be taken from input files
filename_post = name of file containing posterior
filename_prior = name of file containing prior (optional argument)
-r = rescale all plots to the same maximum amplitude. (The default is to plot each marginal on a common amplitude scale, which allows comparison of amplitudes between plots.) If prior/posterior pairs are plotted then they are always on the same scale.
-f = add an annotated frame around each panel. Details of annotation is controlled by options file naplot.cmd
-fn = add an annotated frame around each panel but remove title above each panel. (Because they can get in the way of axis labels).
-n = no annotated frame and no titles.
If options -f or -fn are used and annotation details are not specified in file plot/naplot.cmd then only a simple frame is plotted.
If -r and -f are used then they must be given in the order `-r -f' and not in the order `-f -r'.
Note: Order of marginals in posterior and prior files must be the same. Shading is performed with pen 6. Prior marginal is plotted with pen 4, and reference value is drawn with pen 3. Order of plots can be changed by using input file naplot.cmd (see examples below).
Plotting 2-D marginals of the posterior probability distribution (PPD)
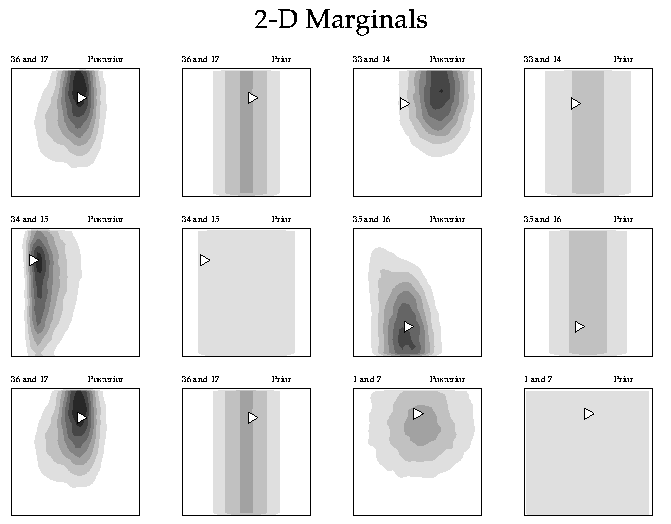
Calling sequence
naplot-x plottype n filename_post (filename_prior) (-r) (-f)
plottype = `2dmarg' to activate.
-r = rescale all contour plots to same maximum amplitude (colour) scale. (The default is to choose colour scale so that all panels are on the same amplitude/colour scale, which allows direct comparison of amplitudes between plots.) If prior/posterior pairs are plotted then they are always plotted on the same scale.
-f = add an annotated frame around each panel. Details of annotation is controlled by options file naplot.cmd
-fn = add an annotated frame around each panel and remove title above each panel. (Can get in the way of axis labels).
-n = no annotated frame and no titles.
All other arguments as for `1dmarg' case.
Note: Order of marginals in posterior and prior files must be the same. Contour shading is performed with pens 11-19 read in from file pal.in. Order of plots can be changed by using input file naplot.cmd (see examples below).
Note that if -r option is used then all panels are amplitude normalized. In this case each panel has a different mapping between PDF value and gray scale, but each maximum amplitude contour will always be black.
The default (no -r) is to choose gray scale so that all panels are on the same scale, hence a single mapping between value and gray scale for all panels. Now only the highest amplitude of all panels plotted will be black and the smallest amplitude of all panels will be white, and this defines the single colour scale for all panels. Note that if different panels are plotted the scale may change.
If prior/posterior pairs are plotted then it only makes sense to plot them on the same amplitude scale and so in this case and the -r option does not affect their relative amplitudes.
So in short the -r option gives maximum contrast within each panel, so you can see the contours, and the default allows true comparison between different panels, which one would normally like to do, darker then means more probability, higher amplitude.
Plotting confidence contours
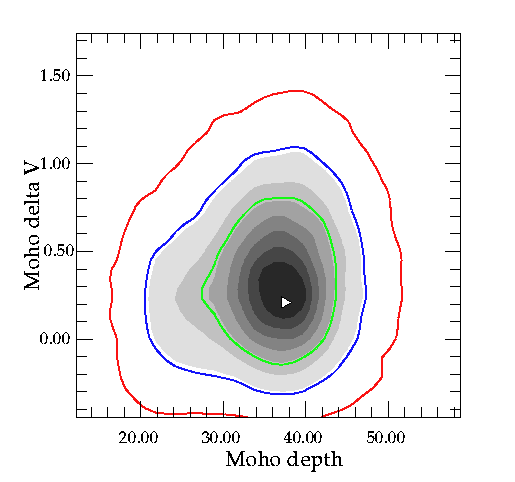
Up to 10 specified confidence contours can be plotted on top of each 2-D marginal distribution. The values of the confidence regions (e.g. 60%,90%,99%), as well as the pen number and the line thickness are read in from the file confidence_cont in the sub-directory plot. (See example file.) The pen colours can be any values, but the corresponding pen colours must be specified in pal.in.
If the file plot/confidence_cont is not present, or has a zero for the number of contours then no confidence contours are plotted. Confidence contours are not plotted on prior PDFs.
Here is an example showing the 60%,90%,99%, using the full annotated frame feature, for the receiver function example problem.
Plotting the posterior Correlation matrix

Calling sequence
naplot-x plottype n filename_post (-t5) (-p)
plottype = `cor' to activate.
n = number of variables to be plotted in correlation matrix
filename_post = name of input file containing posterior (or prior)
-t5 = plot tick marks on matrix every 5 columns and rows. Similarly -tm, where m is any integer.
-p = input file contains prior distribution and so plot labels are amended accordingly (otherwise posterior is assumed).
-e0.2 = plot numerical error in estimated correlation matrix rather than the matrix itself and scale error range by a factor of 0.2. (Similarly for any other value)
Note: shading is performed with pens 41-57 read in from file pal.in. Order of columns and rows can be changed by using input file naplot.cmd (see examples below).
Plotting the posterior Covariance matrix

Calling sequence
naplot-x plottype n filename_post (-t5) (-p) (-e0.2)
plottype = `cov' to activate.
n = number of variables to be plotted in correlation matrix
filename_post = name of input file containing posterior (or prior)
-t5 = plot tick marks on matrix every 5 columns and rows. Similarly -tm, where m is any integer.
-p = input file contains prior distribution and so plot labels are amended accordingly (otherwise posterior is assumed).
-e0.2 = plot numerical error in estimated covariance matrix rather than the matrix itself and scale error range by a factor of 0.2. (Similarly for any other value)
Note: shading is performed with pens 41-57 read in from file pal.in. Order of columns and rows can be changed by using input file naplot.cmd (see examples below).
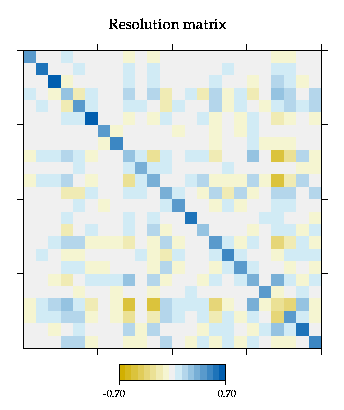
Plotting the Resolution matrix
Calling sequence
naplot-x plottype n filename_post (filename_prior) (-u) (-t5)
plottype = `res' to activate.
filename_post = name of file containing posterior
filename_prior = name of file containing prior (optional). If this argument is not present then the inverse prior covariance matrix is read in from file `plot/inv_prior_cov.matrix'.
-u = plot normal resolution matrix. The default is to plot the non-dimensional resolution matrix.
-t5 = plot tick marks on matrix every 5 columns and rows.
Similarly -tm, where m is any integer.
Note: that off diagonal elements of the normal resolution matrix will depend on dimensional units if more than one variable type is present. The `non-dimensional' resolution matrix is a re-scaled version of the resolution matrix formed by taking the element in the i-th row and j-th column and multiplying by the factor sigma_j/sigma_i, where sigma_i is the square root of the i-th element on the diagonal of the prior covariance matrix.
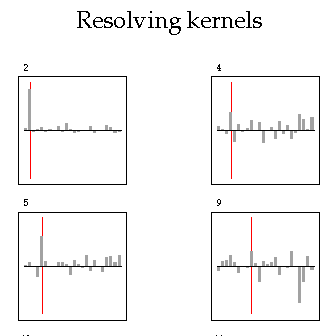
Plotting Resolving kernels
Calling sequence
naplot-x plottype n filename_post (filename_prior) (-c) (-u)
plottype = `reskern' to activate.
filename_post = name of file containing posterior
n = number of variables to be plotted in resolution matrix (maximum = number of variables)
filename_posterior = name of file containing posterior
filename_prior = filename of file containing prior (optional). If this argument is not present then the inverse prior covariance matrix is assumed to be in file `inv_prior_cov.matrix'.
-c = plot columns of the resolution matrix The default is to plot rows.
-u = plot unnormalized resolution matrix. The default is to plot a `non-dimensional' resolution matrix.
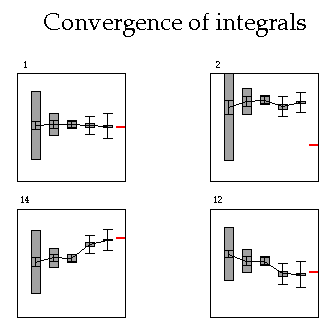
Plotting the convergence of integrals as a function of sample size
Calling sequence
naplot-x plottype n filename_post (-s20)
plottype = `conv' to activate.
n = number of plots to be taken from input files
filename_post = name of file containing posterior
-s20 = scale numerical integration error box by a factor of 20 (makes it more visible). Similarly -sm where m is any integer.
Examples
Here are examples of how to call each plot type with various options. Go to the directory data/utl/naplot where an example prior and posterior can be plotted. (These are both nab.out files.) For more details examine the command file naplot.cmd in the sub-directory plot. More examples can be found in the the file data/utl/naplot/examples.
1-D Marginal distributions:
To plot 1D posterior and prior marginals for first 6 variables,
naplot-x 1dmarg 6 nab_post nab_prior
Plot 24 variables and normalize to same maximum amplitude,
naplot-x 1dmarg 24 nab_post nab_prior -r
Use annotated frame (specified in plot/naplot.cmd),
naplot-x 1dmarg 6 nab_post nab_prior -f
Use zoom feature (specified in plot/naplot.cmd),
cp plot/naplot.cmd_zoom plot/naplot.cmd
naplot-x 1dmarg 6 nab_post nab_prior -f
cp plot/naplot.cmd_full plot/naplot.cmd
Change order of first 4 variables to (1,7,13,19) (Requires entry in naplot.cmd='1 4 1 7 13 19'),
naplot-x 1dmarg 4 nab_post nab_prior -f
If Prior/posterior pairs are plotted they are always area normalized.
2-D Marginal distributions:
Plot 2D posterior and prior marginals for first 6 pairs of variables in input files,
naplot-x 2dmarg 6 nab_post nab_prior
Same as above but normalize all shading to the same maximum amplitude,
naplot-x 2dmarg 6 nab_post nab_prior -r
Plot first 12 2-D marginals, on same shading scale, and add a full annotated frame,
naplot-x 2dmarg 12 nab_post -f
Use zoom feature to plot subsections of first 6 2-D marginals,
cp plot/naplot.cmd_zoom plot/naplot.cmd
naplot-x 2dmarg 6 nab_post -f
cp plot/naplot.cmd_full plot/naplot.cmd
Plot 2D posterior and prior marginals for 5th 2-D marginal in input files (requires line in naplot.cmd = '2 1 5'),
naplot-x 2dmarg 1 nab_post nab_prior
Prior/posterior pairs always have same shading scale.
Correlation and covariance matrix plots:
Plot correlation matrix for first 24 variables, with tick marks every 6th row and column,
naplot-x cor 24 nab_post -t6
36 variables,
naplot-x cor 36 nab_post -t6
Correlation matrix for 12x12 sub-matrix (parameters 1-12)
naplot-x cor 12 nab_post -t6
Correlation matrix for 12x12 sub-matrix (parameters 13-24) (Requires line in naplot.cmd='1 12 12,14,15,...,24)
naplot-x cor 12 nab_post -t6
Plot error in correlation matrix for first 24 variables with error range multiplied by a scale factor of 0.2,
naplot-x cor 24 nab_post -t6 -e0.2
Plot prior correlation matrix for 36 variables,
naplot-x cor 36 nab_prior -t6 -p
Covariance matrix for 12x12 sub-matrix (1-12)
naplot-x cov 12 nab_post -t6
Plot error in Covariance matrix for first 24 variables with a scale factor of 0.2
naplot-x cov 24 nab_post -t6 -e0.2
Note order of variables can be changed in naplot.cmd. All options for `cor' also work for `cov'.
Resolution matrix:
Non-dimensional resolution matrix for first 24 variables (Uses prior covariance read in from file `inv_prior_cov.matrix')
naplot-x res 24 nab_post -t6
As above but resolution matrix,
naplot-x res 24 nab_post -t6 -u
Non-dimensional resolution matrix for first 24 variables with prior covariance taken from nab_prior
naplot-x res 24 nab_post nab_prior -t6
Resolution matrix 12x12 sub-matrix for variables 7-18. (requires line in naplot.cmd = '1 12 7,8,...,18'),
naplot-x res 12 nab_post -t6 -u
Resolution kernels:
Plot rows of the non-dimensional resolution matrix for first 24 variables,
naplot-x reskern 24 nab_post
columns of the non-dimensional resolution matrix for first 12 variables,
naplot-x reskern 12 nab_post -c
Same as above but for variables (19-24) (requires line in naplot.cmd = '1 6 19 20 21 22 23 24')
naplot-x reskern 6 nab_post -c
Plot columns of the resolution matrix for first 6 variables
naplot-x reskern 6 nab_post -c -u
Note order of variables can be changed in naplot.cmd.
Convergence of numerical error in expectation integrals
Plot integration error in expectation integrals for first 24 variables with error boxes ampilified by a 20,
naplot-x conv 24 nab_post -s20
Same as above but for 6 variables (19-24) (requires line in naplot.cmd='1 6 19 20 21 22 23 24'),
naplot-x conv 6 nab_post -s20
References:
The Neighbourhood algorithm for searching a multi-dimensional parameter space for models (points) with `acceptable data fit' is implemented in the program NA-sampler, and is described in the paper:
Geophysical Inversion with a Neighbourhood Algorithm -I. Searching a parameter space, Sambridge, M., Geophys. J. Int., 138, 479-494, 1999. doi:10.1046/j.1365-246X.1999.00876.x
A related paper describes the Neighbourhood re-sampling algorithm (implemented in program NA-bayes) for calculating Bayesian integrals from a finite set of samples produced by the NA algorithm or any other search method (e.g. GA or SA etc.):
Geophysical Inversion with a Neighbourhood Algorithm -II. Appraising the ensemble,Sambridge, M., Geophys. J. Int., 138 ,727-746, 1999. doi:10.1046/j.1365-246x.1999.00900.x
Related sites:
NA homepage - where the latest versions of NA-Bayes and NA-sampler can be accessed.
NA-sampler - the latest version of this manual.
NAD - explanation of direct access nad files.
RFI - Receiver function utility programs and graphics (included with NA-Sampler package.)
NA-Bayes - the manual for the neighbourhood algorithm Bayesian inference package.
splot - graphics program for display of multi-dimensional ensembles.
summary.pdf - A short summary of NA-sampler written for the IASPEI handbook (PDF file)